Upon phage binding to the tip of one of these F pili, disassembly will automatically bring the phage closer to the surface of the bacterium. 
coli partners by a series of F-pilus assembly and disassembly events. coli cells (F +, containing F-pili) appear to trawl the environment in an effort to catch “female” (F −) E. The first stage of the M13 infection is the adsorption process, which takes place through binding of the N2 domain of the G3P coat protein to the tip of a F pilus on the surface of E. Information on the genes and proteins of the M13 phage is listed in Table 1. 
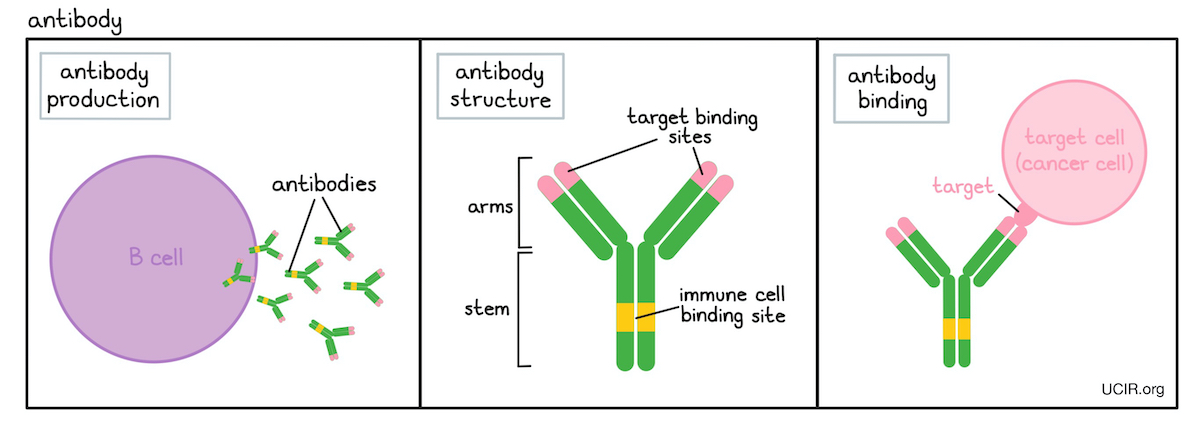
The remaining four coat proteins, G3P, G6P, G7P, and G9P, are each present in approximately five copies. The most abundant of the coat proteins is the capsid protein G8P, which forms an envelope around the chromosome consisting of approximately 2700 protein units ( Figure 1). The M13 phage has a length of 900 nm and a width of 6.5 nm. Five of these proteins are coat proteins, and the remaining six proteins are involved in replication and assembly of the phage. The phage contains a genome of single-stranded DNA (ssDNA) with a length of 6407 bp that consists of nine genes encoding 11 different proteins. Instead, the phage establishes a chronic infection in its host, where it continuously releases new phages. The M13 phage is neither temperate nor lytic. The Ff phages only infect Escherichia coli strains that express the F pilus as the adsorption of the phage to the bacterium requires binding of a phage coat protein to the tip of the F pilus. The M13 phage belongs to a group of filamentous phages collectively referred to as Ff phages. Before giving an in-depth description of the steps involved in phage display experiments, an introduction to the wild-type M13 phage is provided. Therefore, this review will focus on antibody phage display techniques utilizing this specific phage system. However, primarily the M13 phage has been utilized extensively in recent times, and to the best of our knowledge, this is the only phage system that has been explored within toxinology. These different phage systems each have their benefits and drawbacks. Different bacteriophage systems can be utilized for phage display, including the T4, lambda, as well as the filamentous M13 bacteriophage. One approach is phage display selection, which is a robust, easy-to-perform, and inexpensive method by which specific antigen binders are selected from large combinatorial libraries containing billions of antibody fragments.Īs antibody phage display is gaining increasing interest in the field of toxinology, the intention with this review is to provide both basic and more advanced knowledge on the underlying science behind the technology and the lifecycle of the M13 phage.Ĭentral to phage display technology is the biology of the bacteriophage used to display antibodies. To discover and develop antibodies, different techniques can be harnessed. Moreover, these therapeutic proteins could be manufactured cost-competitively using modern cell cultivation methods employed for large scale production. One promising approach seems to be the use of human IgG antibodies and/or camelid antibody fragments, as these molecules can be used to develop recombinant antivenoms with high efficacy and safety due to their compatibility with the human immune system. Several different avenues aimed at bringing innovation into the field of snakebite antivenoms have been pursued, including medicinal chemistry approaches, novel immunization techniques, and the use of biotechnological strategies. This creates renewed hope for snakebite victims worldwide and could potentially lead to a mobilization of scientific efforts toward the development of novel snakebite envenoming therapies. 
With the recent inclusion of snakebite envenoming on the World Health Organization’s list of Neglected Tropical Diseases, focus on both prevention and treatment of this infliction has increased. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |



 RSS Feed
RSS Feed
